Welcome to tRFTar!
Introduction
tRNA-derived fragments (tRFs), which by definition are cleaved from tRNAs, comprise a novel class of regulatory small non-coding RNAs.
Recent evidence has revealed that tRFs can be loaded onto Argonaute (AGO) family proteins to perform post-transcriptional regulations via
substantial tRF-target gene interactions (TGIs). Here we performed a systemic computational screening of potential AGO-mediated TGIs by a
re-analysis of 146 crosslinking-immunoprecipitation and high-throughput sequencing (CLIP-seq) datasets in which 920,690 TGIs between
12,102 tRFs and 5,688 target genes were identified. Furthermore, filtering TGIs with consistent co-expression with target genes
results in a set of regulatory TGIs that contains 25,281 tRF-target pairs.
All the CLIP-derived TGIs are incorporated in this platform and there are various functions such as custom searching, co-expressed TGI filtering,
genome browser, TGI-based tRF functional enrichment analysis and so on to help researchers to investigate the functions of tRFs (See 'Tutorial'
for detail).
Citation
Zhou Y, Peng H, Cui Q, Zhou Y. tRFTar: Prediction of tRF-target gene interactions via systemic re-analysis of Argonaute CLIP-seq datasets. Methods. 2021 Mar;187:57-67.
For the convenience of downstream researches, tRFTar related data (.csv format) can be download below:
Co-expression profile: Download
TGIs presented as duplexes (all tRFs): Download
TGIs presented as duplexes (top highly expressed tRFs): Download
TGIs presented as tRF-element pairs (all tRFs): Download
TGIs presented as tRF-element pairs (top highly expressed tRFs): Download
KIRC related tRFs: Download
tRF-Gene Interaction(TGI) Search and tRF Functional Enrichment Analysis are the main
functions provided by tRFTar.
TGI Search
TGIs collected in this platform were dug out from crosslinking-immunoprecipitation and high-throughput sequencing (CLIP-seq) datasets, please read the tRFTar
paper for more detail. All the CLIP-derived TGIs can be roughly browsed in the Browse Page (enabled by clicking the 'Browse' item of the navigation bar).
Moreover, users also can query specific interested TGIs by quick search or advanced search.
Quick Search
1. Input an interested tRF MINTBase ID/sequence or gene symbol in the Quick Search Input Box of the navigation bar, and then press the 'Enter' key to launch the
search task.

2. All CLIP-derived TGIs which involve the inputted tRF or gene will be listed in the Search Result Page.
Advanced Search
1. Click the 'Search' item of the navigation bar to enable the Advanced Search Page.
2. Input the tRF MINTBase ID/sequence list and the gene symbol list, and then the search result will be restricted to TGIs between these inputted tRFs and genes.
Note that if the tRF list is left blank, all the tRFs will be considered without restriction, as well as the gene list.
3. Set advanced options according to requirements and click the 'Search' button to launch the search task. In the current version, users can search specific
CLIP-derived TGIs by options of specifying tRF classes and target element classes, restricting tRFs to be highly-expressed and restricting tRF-gene pairs to
be co-expressed.
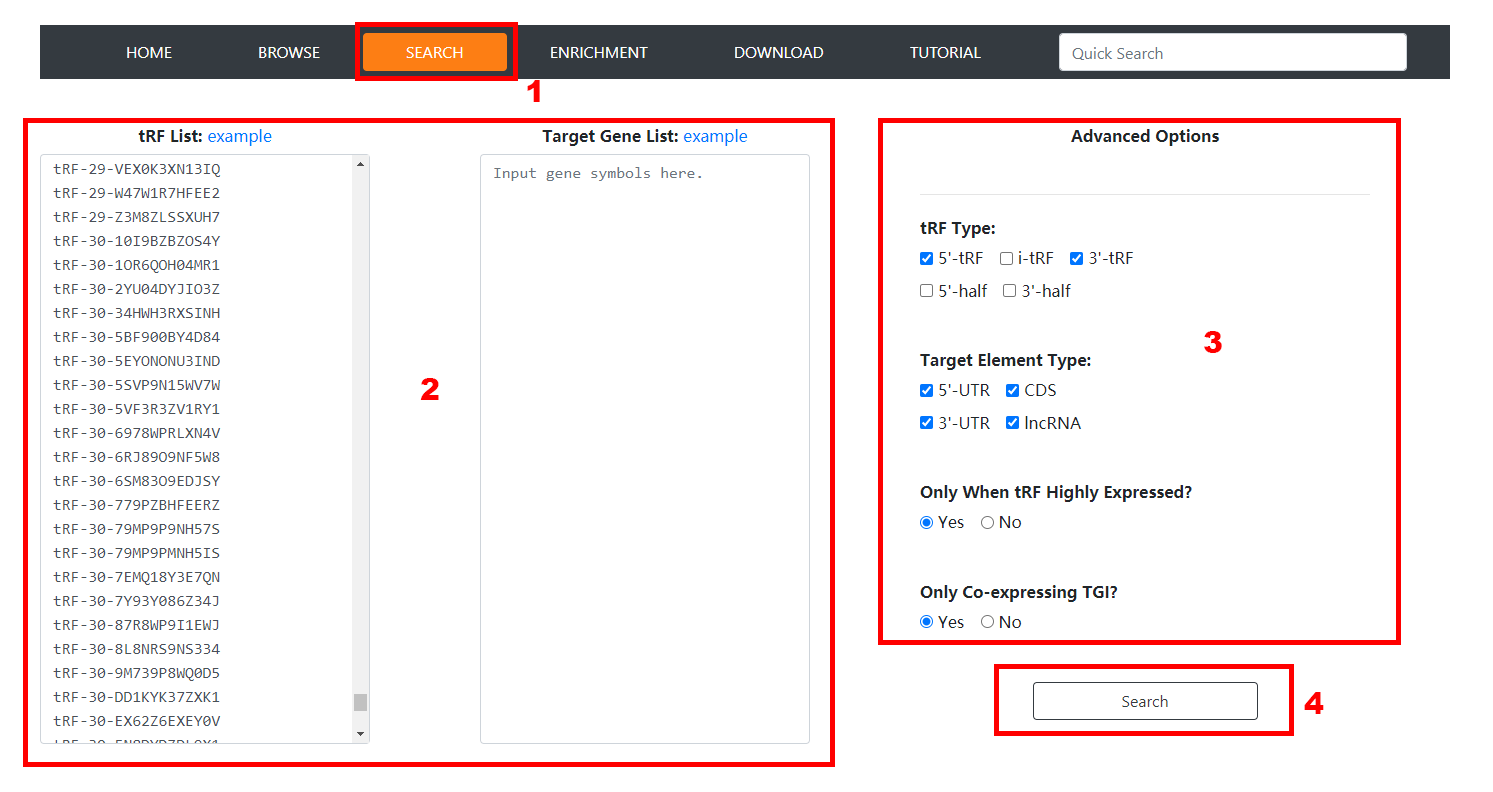
4. All CLIP-derived TGIs matching the query criteria will be listed in the Search Result Page.
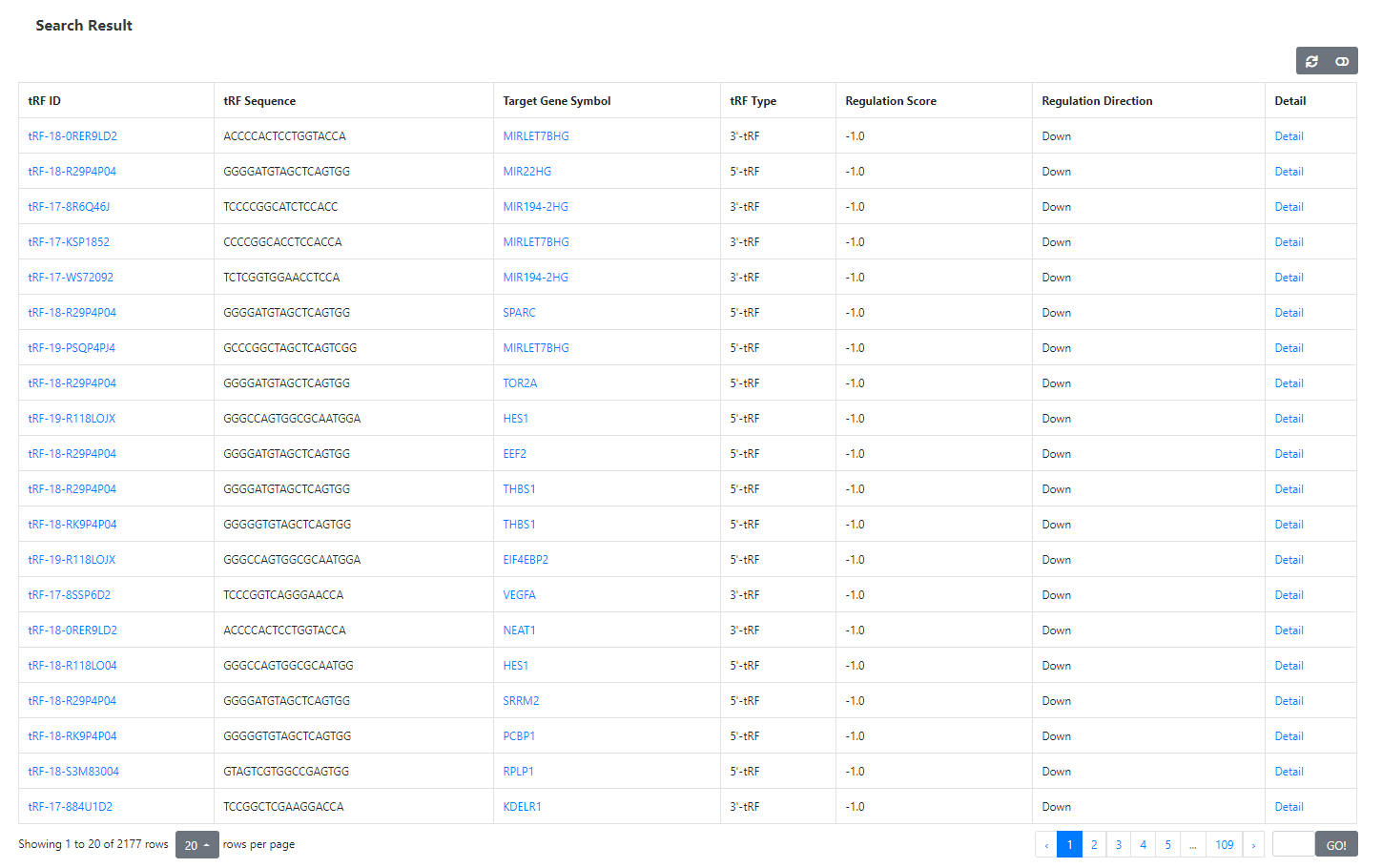
The Search Result Page
The Search Result Page lists the basic information of TGIs matching the query criteria, including tRF ID, tRF sequence, tRF type, target gene symbol and
regulation score (an indicator introduced in the tRFTar paper). Users can browse the detailed information of a certain TGI by clicking the 'Detail'
link in the corresponding entry.
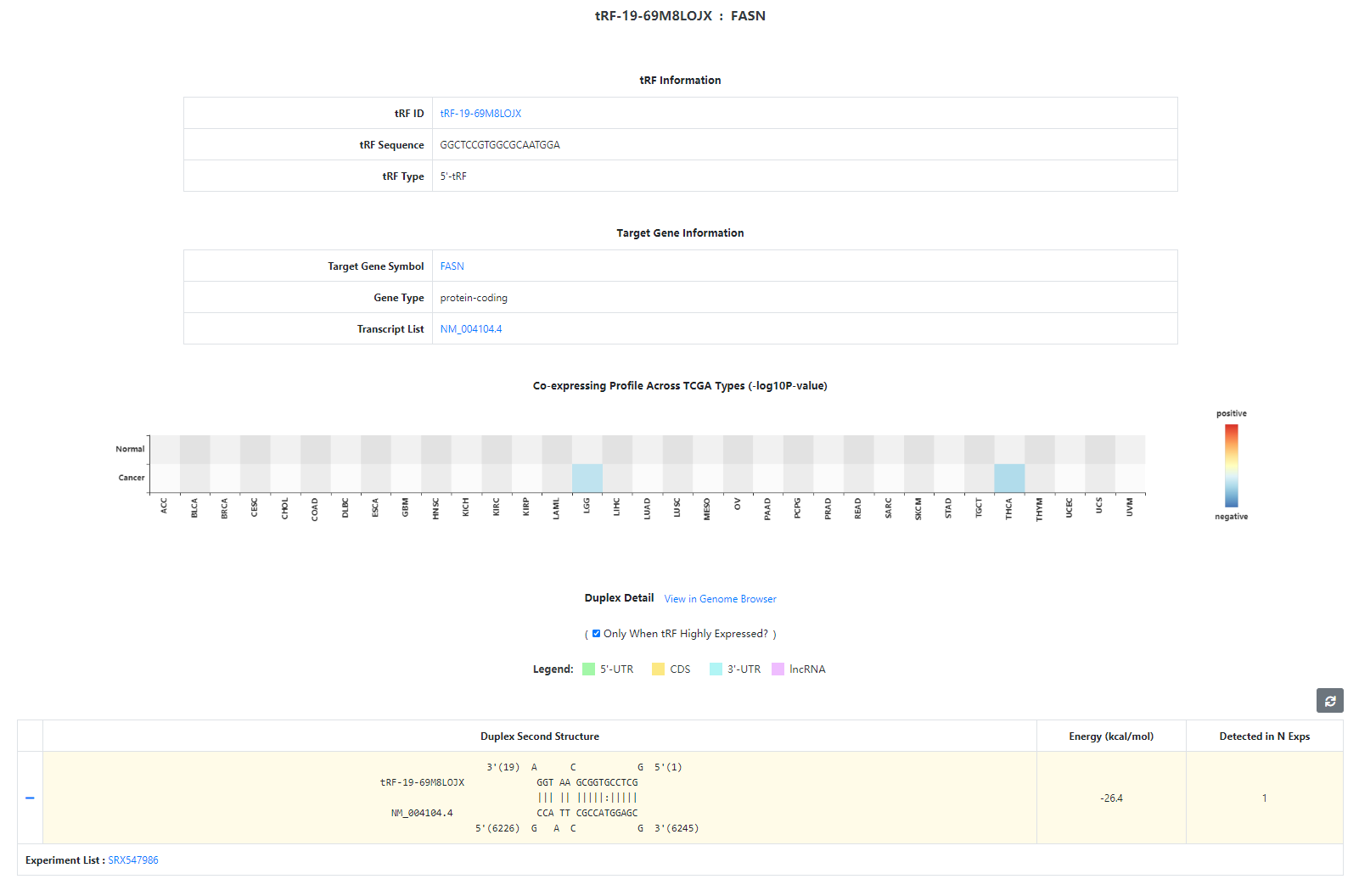
The TGI Detail Page
The TGI Detail Page shows the detailed information of a certain TGI, including the basic information of tRF and target gene, the co-expression profile
and the detailed information of tRF-target region duplexes. Note that the co-expression profile is visualized by signs of Spearman correlation coefficients
and -log10-transformed p-values (only entries whose p-value ≤ 0.01 are retained) across TCGA types. For the detailed information of a
certain duplex, besides the conspicuously displayed second structure and minimum free energy, links of CLIP datasets where the duplex is detected can be
expanded by clicking the left side '+' button. Moreover, tRF target sites can be viewed in the genome browser by accessing the 'View in Genome Browser' link.
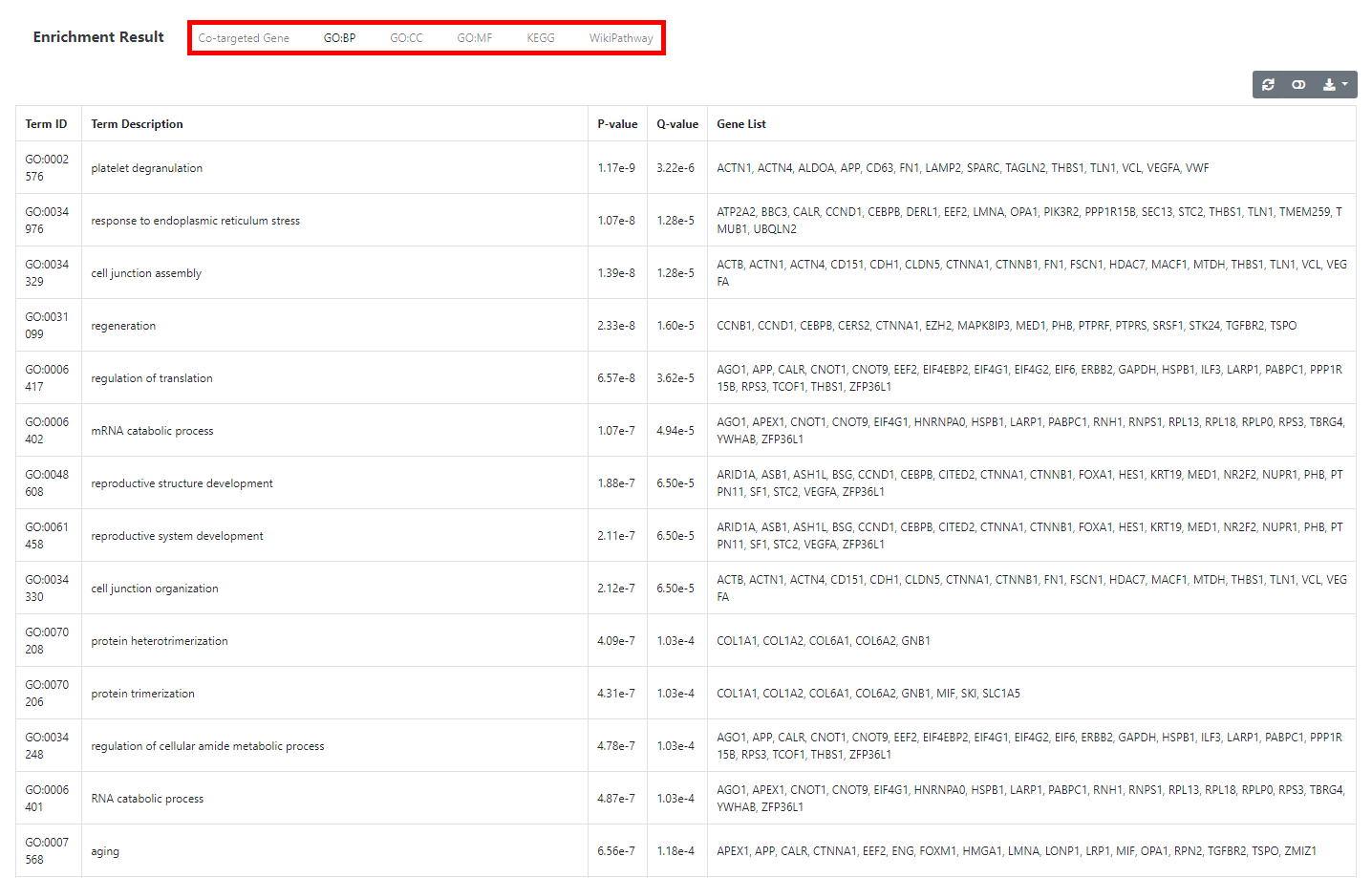
tRF Functional Enrichment Analysis
In this module, functions of inputted tRFs can be inferred from the function annotations of their target genes. Specifically, a tRF list is expected to be inputted
at first, then genes co-targeted by N (a user-defined integer) tRFs of the list will be found out. Finally, co-targeted genes will be submitted to the enrichment
analyses against optional well-known functional gene set annotations such as Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) and WikiPathway
so that related functions of inputted tRFs can be deduced. Operation steps are described below:
1. Click the 'Enrichment' item of the navigation bar to enable the tRF Functional Enrichment Analysis Page.
2. Input the tRF MINTBase ID/sequence list.
3. Set advanced options according to requirements and click the 'Analysis' button to launch thr enrichment analysis task. In the current version, options of specifying
functional gene set annotations, restricting tRFs to be highly-expressed, restricting tRF-gene pairs to be co-expressed and setting the targeting tRF number N are
provided.
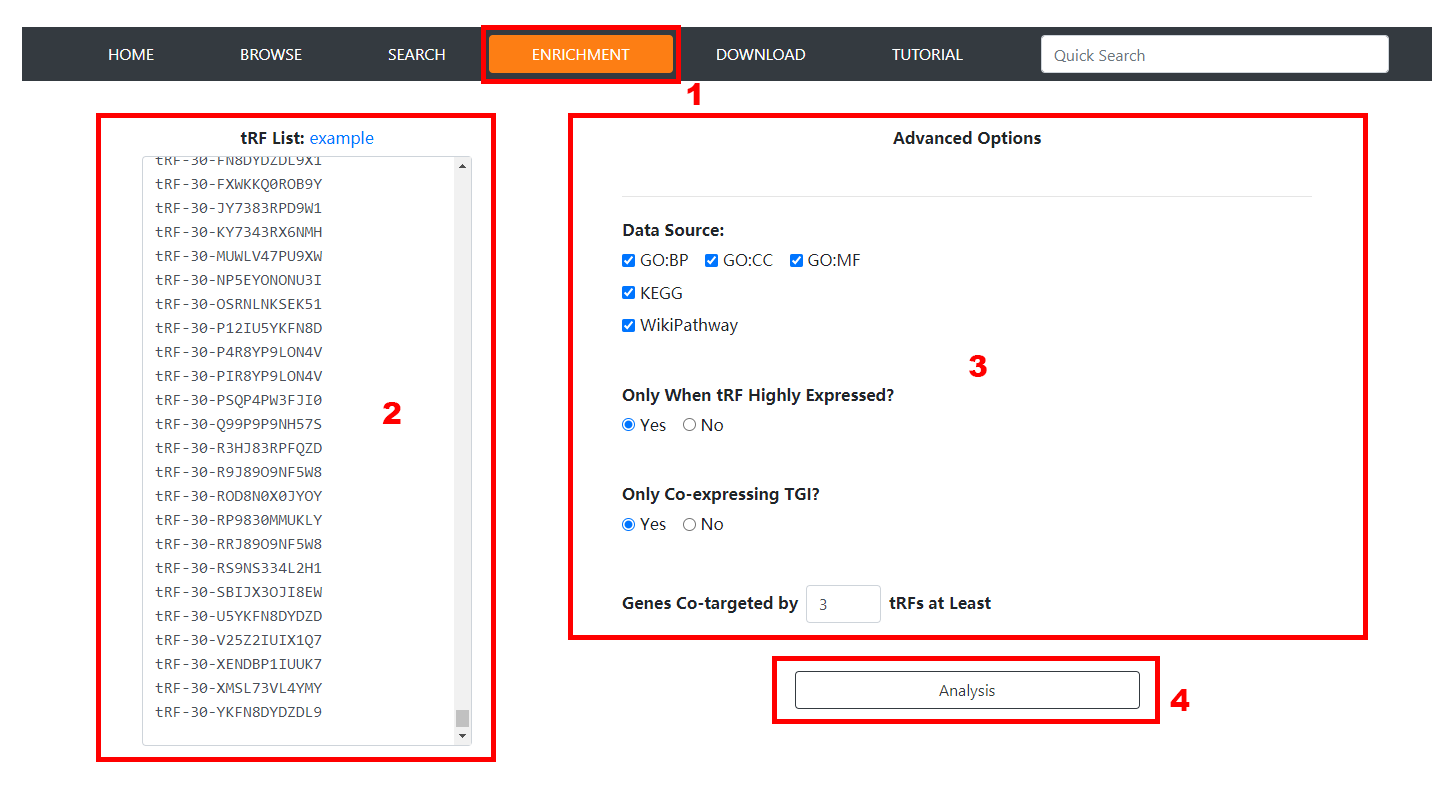
4. Co-targeted genes and enrichment results will be listed in the Enrichment Result Page, which can be mutually shown by clicking corresponding navigation items.